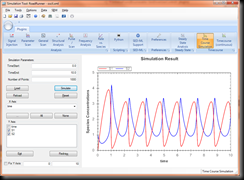
With the release 2.8.1 of the Systems Biology Workbench the Simulation Tool now has a plug-in system based on MEF. This enabled me to enhance the simulation tool by both new layouts (changing the look and feel) as well as include new Simulation Experiments (like the Signal Injection plug-in). But now that we have this new plug-in API it also allows Y O U to write for it. And I’m excited to see Y O U R contributions. (I’m preparing a online repository for plug-ins). So how do you write these plug-ins? In this series I show you how to set up the IDE, so that you can start writing plug-ins, and then give a couple of examples of simple plug-ins one could write.
There are three ways that I envision people would write plug-ins:
Visual Studio
Visual Studio is my tool of choice when it comes to programming. Luckily most universities have contracts with microsoft, so ask your IT department for a copy (ours had 2010 Ultimate!). If they don’t you still have the option of getting a full version at dreamspark:
https://www.dreamspark.com/
Or of course the express editions will work:
http://www.microsoft.com/express/Default.aspx.
In the following I use screenshots from Visual Studio 2010, but the same will apply to Visual Studio 2008 and Express editions.
In order to ease the process of writing plug-ins I have created Visual Studio Templates. You can download them from:
http://sourceforge.net/downloads/sbwsimtool/visual-studio-templates/v1
simply download the Zip files and place them into:
C:\Users\fbergmann\Documents\Visual Studio 2010\Templates\ProjectTemplates\Visual C#
(just replace fbergmann with whatever your user name is). The same templates also work for Visual Studio 2008 where you place them into:
C:\Users\fbergmann\Documents\Visual Studio 2008\Templates\ProjectTemplates\Visual C#
Once this is done start Visual Studio and create a new project. Now in the template wizard you can simply select the SimulationToolPlugin template:
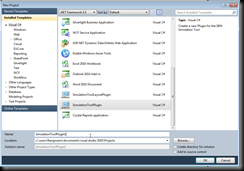
Once you confirm the new project will be created and loaded. The code already implements a simple plug-in. So let us set up the properties, so that we can run it. For this go to
Project\SimulationTool1Plugin Properties
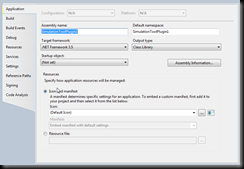
Here you want to change the ‘Assembly Information’ to provide a short description of your plug-in an set your Copyright information. Next you might want to change the Default Namespace and the Assembly Name. My recommendation is to use the same name for both. The Assembly Name controls what the name of the resulting .DLL will be.
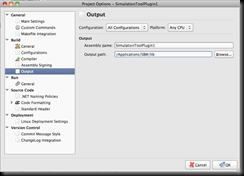
Next let us flip to the Build tab:
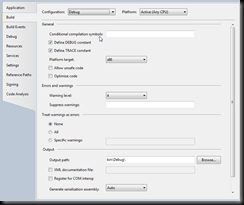
The key elements here are: The platform target: make sure to select x86 (as the simulation tool uses native libraries that are compiled for 32bit only). The output path:
you want to set this one *for now* to the directory of the Simulation Tool:
C:\Program Files (x86)\KGI\SBW\roadrunner
on 64-bit systems and:
C:\Program Files\KGI\SBW\roadrunner
on 32 bit systems.
Finally, let us set up how to start the debug session:
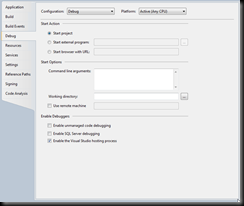
we want to start an external program:
C:\Program Files (x86)\KGI\SBW\roadrunner\SimDriverNET.exe or
C:\Program Files\KGI\SBW\roadrunner\SimDriverNET.exe
and we want to set the working directory to:
C:\Program Files (x86)\KGI\SBW\roadrunner or
C:\Program Files\KGI\SBW\roadrunner
(One note for express users: you won’t be able to start an external program in the express edition … I suggest you choose a Post Build Event to start the SimDriverNET executable by adding:
cd C:\Program Files\KGI\SBW\roadrunner && SimDriverNET.exe
as post build event now after every build you see the simdriver loaded with the new plugin, alas no debugging support.)
THAT IS IT … now we are ready to test our plug-in! close the property tab and press RUN. Our new plug-in is in an unnamed category so you can easily find it:
And when you load a model you can perform time course scans by entering values into it. Here a screenshot for the ‘oscli.xml’ file:
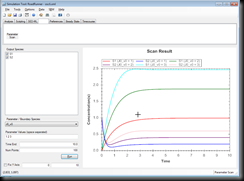
This is it you are all set up and ready to play with the plug-in.